Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) causes coronavirus disease 2019 (COVID-19) and has reached pandemic status since the first cases were detected in Wuhan, Hubei province, China, in late 2019. Despite its rapid worldwide spread, many unknown aspects are to be discovered regarding epidemiological and clinical patterns of COVID-19, especially among children. SARS-CoV-2 infection involves mainly middle-aged and elderly individuals, and the mortality rates are highest in compromised older adults with underlying diseases. Constantly, the reported proportion of children with SARS-CoV-2 infection has been low. The main transmission pathways are respiratory droplets, contact, and potentially fecal-oral. Despite the clinical evolution of adults, the paediatric patients with COVID-19 have relatively milder symptoms, and this difference is not fully understood. Some hypotheses suggest that the amount of viral loads (or the duration of virus-shedding period) is less important or shorter in children. The most important finding is that we now have clear evidence that children are susceptible to SARS-CoV-2 infection, but mostly there is not any serious illness, raising the possibility that children could be viral transmiters and urging investigation regarding their role in the epidemiologic chain.
Patogenia infecţiei cu SARS-CoV-2 la om
Pathogenesis of SARS-CoV-2 infection in humans
First published: 23 mai 2020
Editorial Group: MEDICHUB MEDIA
DOI: 10.26416/Pedi.58.2.2020.3572
Abstract
Rezumat
SARS-CoV-2, un nou tip de coronavirus, produce boala denumită COVID-19, care are caracteristici pandemice, extinzându-se cu rapiditate încă de la depistarea primelor cazuri, în Wuhan, provincia Hubei din China, la finalul anului 2019. Deşi a implicat o arie populaţională extinsă, prin nivelul înalt de contagiozitate, există încă multe aspecte necunoscute ce ţin de caracteristicile epidemiologice şi patogenice ale COVID-19, mai ales la vârsta pediatrică. Nivelul de agresivitate al infecţiei cu SARS-CoV-2 este proporţional cu vârsta, iar mortalitatea este importantă la cei cu boli cronice asociate, spre deosebire de populaţia pediatrică, unde ratele de îmbolnăvire au fost constant raportate ca fiind scăzute. Calea de transmitere majoră este reprezentată de picăturile respiratorii, urmată de contactul direct şi, potenţial, calea fecal-orală. În contrast cu evoluţia de la adulţi, copiii dezvoltă forme uşoare de boală, deşi mecanismele nu sunt încă în totalitate cunoscute. Unele ipoteze au sugerat faptul că nivelul încărcăturii virale şi perioada de replicare sunt mai reduse la copii. Astăzi ştim cu siguranţă faptul că populaţia pediatrică este susceptibilă la infecţia cu SARS-CoV-2, dezvoltând forme uşoare şi moderate de boală, devenind astfel un important transmiţător viral, ceea ce impune investigaţii epidemiologice extinse.
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) causes coronavirus disease 2019 (COVID-19) and has reached pandemic status since the first cases were detected in Wuhan, Hubei province, China, in late 2019.
Coronaviruses are known for causing mild upper respiratory tract infections, but in their family, previously, Severe Acute Respiratory Coronavirus (SARS-CoV) and Middle East Respiratory Syndrome Coronavirus (MERS-CoV) have been implicated in fatal or severe illnesses. It is well known that infants and young children typically have a high rate of hospital admission for respiratory tract infections, most commonly caused by viral infections with germs such as the respiratory syncytial virus, rhinovirus, human metapneumovirus and influenza virus, due to immaturity of the respiratory tract and the immune system particularities(1). Nevertheless, it is surprising the pattern of epidemiology and clinical evolution of COVID-19 in paediatric patients. So far, a remarkable characteristic is that only a small number of patients confirmed with SARS-CoV-2 infection are children(2).
Despite its rapid worldwide spread, many unknown aspects are to be discovered regarding the epidemiological and clinical patterns of COVID-19, especially among children. There is no current recommendation regarding specific antiviral medicine, although several researches have been conducted to evaluate treatment options(3). A special interest in the field was attributed to host immune system, known to have a vital role against SARS-CoV and MERS infections. The main characteristic of this activation is the change in peripheral blood T lymphocyte subsets, thus considered a key for understanding the evolution and treatment of the disease(4).
Basic virology
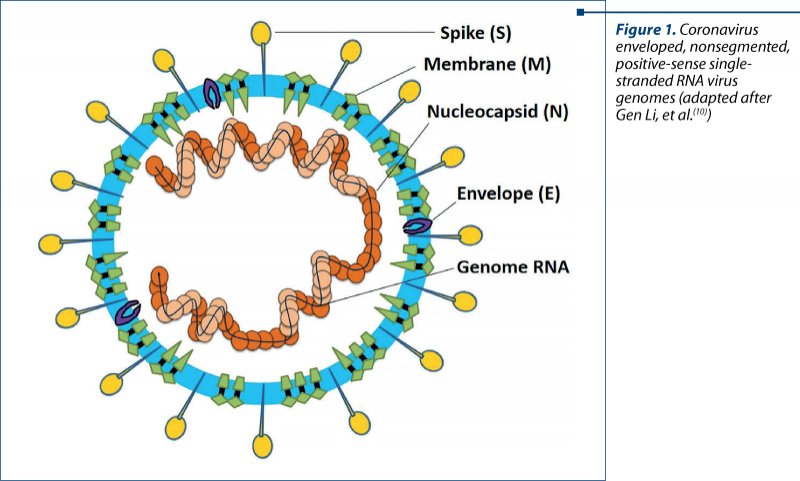
Coronaviruses (CoVs) are single-stranded RNA viruses that infect a broad range of vertebrates; bats are a major host, as well as birds, mammals and humans(5). The previously mentioned SARS-CoV and MERS-CoV are phylogenetically distinct from the common human CoVs, originated mainly in bats and having human as an intermediate host(6,7). Their evolution was more severe in humans, with stronger virulence, rapidly passing from person to person, producing intense alveolar damage and leading to gradual respiratory failure. Regarding SARS-CoV-2 as a component of this family, studies found out that it shares 88% sequence with two bat-derived SARS-like CoV, suggesting it had originated in bats, and the molecular sequencing identified 79.5% overlap between SARS-CoV-2 and SARS-CoV(8). As other coronaviruses, SARS-CoV-2 genome encodes four major structural proteins: the spike (S) protein, nucleocapsid (N) protein, membrane (M) protein, and the envelope (E) protein(9). Out of these, entrance in the target cell is enabled by the S protein, due to its short intracellular tail, transmembrane anchor and its large ectodomain (consisting of S1 subunit and a membrane-fusing S2 subunit)(7). Figure 1 synthetically illustrates the structural components of the new coronavirus.
Pathogenesis
SARS-CoV-2 infection involves mainly middle-aged and elderly individuals, and the mortality rates are highest in compromised older adults with underlying diseases. Constantly, the reported proportion of children with SARS-CoV-2 infection has been low. The paediatric cases of coronavirus disease 2019, caused by severe acute respiratory syndrome coronavirus 2, have been reported. In the United States of America, 2% of confirmed cases of COVID-19 were among persons aged <18 years old, and in China 2.2% of confirmed cases of COVID-19 were among persons aged <19 years old(11). In Italy, 1.2% of COVID-19 cases were among children aged <18 years old(12). Another major European cluster, in Spain, revealed that 0.8% of confirmed cases of COVID-19 were among persons aged <18 years old(13).
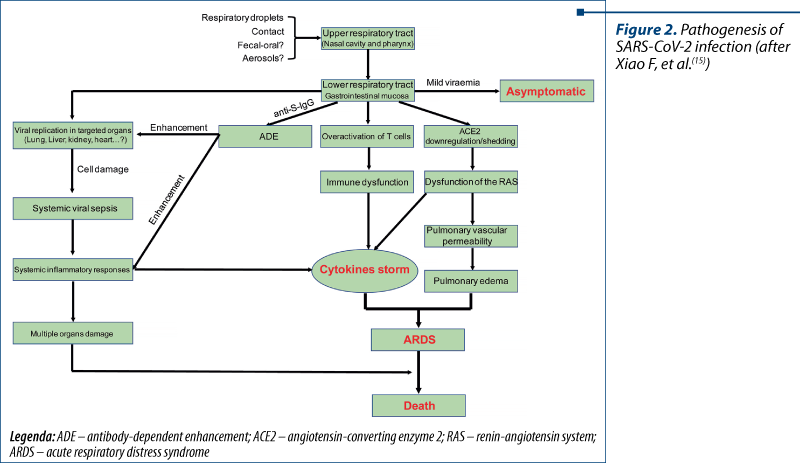
The main transmission pathways are respiratory droplets (the direct pathway), the contact with contaminated surfaces and objects (indirect parthway), and potentially fecal-oral(14). The replication begins in the mucosal epithelium of upper respiratory tract (nasal cavity and pharynx), and is continued in the lower respiratory tract or gastrointestinal mucosa(15), resulting in mild viremia. The structural similarity between SARS-CoV, MERS-CoV and SARS-CoV-2 gives us a lot of information on the pathogenesis of COVID-19, based on common mechanisms. The entire pathogenesis flow is shown in Figure 2.
Entrance and replication. The data available so far suggested that the virus enters the human alveolar epithelial cells using receptors on the cell surface and begins its replication using cellular components(16). The envelope spike glycoprotein binds to its cellular receptor which is angiotensin-converting enzyme 2 (ACE2)(7). Such receptors are found in many other tissues beside respiratory epithelium, like heart, esophagus, stomach, bladder, ileum and kidney, making them vulnerable to SARS-CoV-2(17). Entering in the cell enables the virus to release viral RNA genome into the cytoplasm and then this is translated into two polyproteins and structural proteins; the process is followed by genome replication(18). The new glycoproteins binds the membrane of the endoplasmic reticulum or Golgi in order to form a nucleocapsid and vesicles containing viral particles fuse the plasma membrane and release the virus(19).
Antigen presentation. Antigenic peptides of SARS-CoV-2 are presented to the antigen presentation cells (APC) by the major histocompatibility complex (MHC or human leukocyte antigen [HLA] in humans); thus, they are perfectly recognized by the virus-specific cytotoxic T lymphocytes (CTLs). This is a supposition based on SARS-CoV and MERS-CoV studies on MHC I molecules(20). There is a large HLA polymorphism correlated to infection susceptibility.
Humoral and cellular immunity. The recognition of the antigen stimulates humoral and cellular immunity mediated by virus-specific B and T cells, leading to antibodies production like in other viral infections. In SARS infection, specific IgM antibodies disappear at the end of week 12 and IgG antibody last for a longer period, indicating their protective role(21). Regarding cellular immunity, the peripheral blood of SARS-CoV-2-infected patients is defective of CD4+ and CD8+ T cells in the acute phase. In the absence of antigen, these memory T cells can persist up to four years in some SARS-CoV recovered individuals, inducing T cell proliferation, DTH response and production of IFN-g in case of reinfection(22).
Cytokine storm is considered to be one of the main mechanisms for severe pulmonary infection in COVID-19 patients, and is characterized by uncontrolled systemic inflammatory response due to release of large amounts of proinflammatory cytokines (IFN-a, IFN-g, IL-1b, IL-6, IL-12, IL-18, IL-33, TNF-a, TGFb) and chemokines (CCL2, CCL3, CCL5, CXCL8, CXCL9, CXCL10) by immune effector cells in SARS-CoV infection(22).
A particular viral surviving mechanism, namely coronavirus immune evasion, enables some protective pathways against host immunity: production of double-membrane vesicles that lack pattern recognition receptors (PRRs) and then replicate in these vesicles, thereby avoiding the host detection of their dsRNA, membrane proteins of MERS-CoV inhibit nuclear transport of IFN regulatory factor 3 (IRF3) and activation of IFN b promoter, the IFN-I pathway (protective role against infection) is inhibited in infected mice(23).
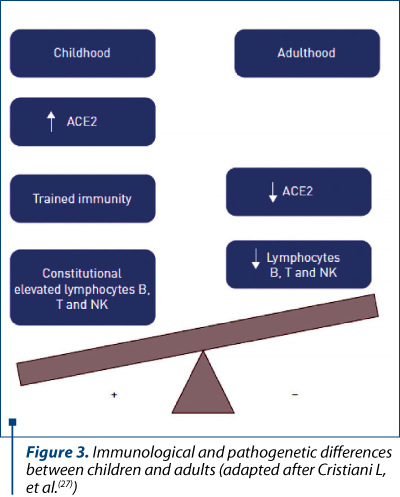
Despite the clinical evolution in adults, paediatric COVID-19 patients have relatively milder symptoms and this difference is not fully understood. Some hypotheses suggest that the amount of viral loads (or the duration of virus-shedding period) is less important or shorter in children(24). Another possibility is that the expression level of ACE2 may differ between ages because researchers showed that ACE2 is more abundant on mature well-differentiated ciliated epithelial cells(25). Additionally, gender could also affect ACE2 expression, considering that ACE2 gene is located on the X-chromosome and the levels of circulating ACE2 are lower in women than in men(26). A very recent publication illustrated the immunological and pathogenetic differences between children and adults consisting in elevated ACE2 expression and lymphocyte count in younger ages; in the meantime, children follow immunization schedules and face viral infections as a potential booster for their innate and adaptive immunity (Figure 3)(27).
Also, studies have suggested that children have a qualitatively different response to the virus than adults, because antigen stimulation and thymic involution determine naive T cells to become effector memory T cells, effector T cells and central memory T cells(26). Another explanation is that children have a simultaneous presence of other viruses in the mucosa lungs and airways, thus SARS-CoV-2 compete with them and its own replication is restricted(28).
Conclusions
The most important finding is that we now have clear evidence that children are susceptible to SARS-CoV-2 infection, but mostly there is not any serious illness, raising the possibility that children could be viral transmiters and urging investigation regarding their role in the epidemiologic chain.
Conflicts of interests: The authors declare no conflict of interests.
Bibliografie
- Hong L, Luo Y. Respiratory viral infections in infants: causes, clinical symptoms, virology, and immunology. Clin Microbiol Rev. 2010; 23(1), 74–98.
- Chan JF, Yuan S, Kok KH, To KK, Chu H, Yang J, et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020; 395(10223), 514-523.
- Zhao J, Alshukairi AN, Baharoon SA, Ahmed WA, Bokhari AA, Nehdi AM, Layqah LA, Alghamdi MG, Al Gethamy MM, Dada AM, Khalid I, Boujelal M, Al Johani SM, Vogel L, Subbarao K, Mangalam A, Wu C, Ten Eyck P, Perlman S, Zhao J. Recovery from the Middle East respiratory syndrome is associated with antibody and T-cell responses. Sci Imm. 2017; 2(14), eaan 5393.
- Lin L, Lu L, Cao W, Li T. Hypothesis for potential pathogenesis of SARS-CoV-2 infection – a review of immune changes in patients with viral pneumonia. Emerging Microbes & Infections. 2020; 9(1), 727-732.
- Fehr AR, Perlman S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol. 2015; 1282, 1-23.
- Gu J, Korteweg C. Pathology and pathogenesis of severe acute respiratory syndrome. Am J Pathol. 2007; 170(4), 1136-1147.
- Zhou P, Yang XL, Wang XG, HU b, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chean HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XZ, Zhoa K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020; 579, 270-273.
- Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020; 395(10224),565-574.
- Schoeman D, Fielding BC. Coronavirus envelope protein: current knowledge. Virol J. 2019; 16(1): 69.
- Li G, Fan Y, Lai Y, Han T, Li Z, Zhou P, Pan P, Wang W, Hu D, Liu X, Zhang Q, Wu J. Coronavirus infections and immune responses. J Med Virol. 2020; 1–9.
- Wu Z, McGoogan JM. Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72314 Cases From the Chinese Center for Disease Control and Prevention. JAMA. 2020; 323(13), 1239-1242.
- Livingston E, Bucher K. Coronavirus Disease 2019 (COVID-19) in Italy. JAMA. doi: 10.1001/jama.2020.4344.
- Tagarro A, Epalza C, Santos M, et al. Screening and Severity of Coronavirus Disease 2019 (COVID-19) in Children in Madrid, Spain. JAMA Pediatr. 2020; doi: 10.1001.
- General Office of National Health Commission; General Officece of National Administration of Traditional Chinese Medicine. Diagnostic and treatment protocol for Novel Coronavirus Pneumonia; (Trial version 6). Available online: http://www.nhc.gov.cn/yzygj/s7653p/202002/8334a8326dd94d329df351d7da8aefc2.shtml
- Xiao F, Tang M, Zheng X, Li C. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology. 2020; 158(6), 1831-1833.
- Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell. 2020; 181(2), 271-280.
- Zou X, Chen K, Zou J, Han P, Hao J, Han Z. The single-cell RNA-seq data analysis on the receptor ACE2 expression reveals the potential risk of different human organs vulnerable to Wuhan 2019-nCoV infection. Front Med. 2020; 1-8.
- Perlman S, Netland J. Netland Coronaviruses post-SARS: update on replication and pathogenesis. Nat Rev Microbiol. 2009; 7(6), 439-450.
- de Wit E, van Doremalen N, Falzarano D, Munster VJ. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016; 14(8), 523-534.
- Liu J, Wu P, GaoF, Qi J, Kawana-Tachikawa A, Xie J, Vavricka CJ, Iwamoto A, Li T, Gao GF. Novel immunodominant peptide presentation strategy: a featured HLA-A*2402-restricted cytotoxic T-lymphocyte epitope stabilized by intrachain hydrogen bonds from severe acute respiratory syndrome coronavirus nucleocapsid protein. J Virol. 2010; 84(22), 11849-11857.
- Li G, Chen X, Xu A. Profile of specific antibodies to the SARS-associated coronavirus. N Engl J Med. 2003; 349(5), 508-509.
- Fan YY, Huang ZT, Li L, Wu MH, Yu T, Koup RA, Bailer RT, Wu CY. Characterization of SARS-CoV-specific memory T cells from recovered individuals 4 years after infection. Arch Virol. 2009; 154(7), 1093-1099.
- Li X, Geng M, Peng Y, Meng L, Shemin L. Molecular immune pathogenesis and diagnosis of COVID-19. Journal of Pharmaceutical Analysis. 2020; 10, 102-108.
- Liu Y, Yan LM, Wan L, Xiang TX, Le A, Liu JM, Peiris M, Poon LLM, Zhang W. Viral dynamics in mild and severe cases of COVID-19. Lancet Infect Dis. 2020; S1473-3099(20), 30232-2.
- Jia HP, Look DC, Shi L, Hickey M, Pewe L, Netland J, Farzan M, Wohlford-Lenane C, Perlman S, McCray PB Jr. ACE2 receptor expression and severe acute respiratory syndrome coronavirus infection depend on differentiation of human airway epithelia. J Vir. 2005; 79(23), 14614–14621.
- Patel SK, Velkoska E, Burrell LM. Emerging markers in cardiovascular disease: where does angiotensin-converting enzyme 2 fit in? Clin Exp Pharmacol Physiol. 2013; 40(8), 551–559.
- Saule P, Trauet J, Dutriez V, Lekeux V, Dessaint JP, Labalette M. Accumulation of memory T cells from childhood to old age: central and effector memory cells in CD4(+) versus effector memory and terminally differentiated memory cells in CD8(+) compartment. Mech Ageing Dev. 2006; 127(3), 274–281.
- Cristiani L, Mancino E, Matera L, Nenna R, Pierangeli A, Scagnolari C, Midull F. Will children reveal their secret? The coronavirus dilemma. Eur Respir J. 2020; 55, 2000749.
- Nickbakhsh S, Mair C, Matthews L, Reeve R, Johnson PCD, Thorburn F, von Wissmann B, Reynolds A, McMenamin J, Gunson RN, Murcia PR. Virus-virus interactions impact the population dynamics of influenza and the common cold. Proc Natl Acad Sci USA. 2019; 116(52), 27142-27150.
Articole din ediţiile anterioare
Provocări în pandemie: pneumotoraxul la copil
Pandemia de COVID-19 cu care încă ne confruntăm continuă să aducă noi provocări în gestionarea multor boli cronice care îşi modifică evoluţia.
Acute liver failure associated with recent SARS-CoV-2 infection in a pediatric patient – case report
Insuficienţa hepatică acută (IHA) la copil este o patologie rară, dar severă, caracterizată prin coagulopatie şi modificări de laborator sugestive ...
Schimbare de paradigmă în MIS-C – cazul unei furtuni perfecte
Infarctul renal acut la copii şi adolescenţi este un eveniment rar, asociat de obicei cu anomalii cardiace, cu status hipercoagulant, boli cu ...
Elemente actuale privind fiziopatologia cirozei hepatice la copil
Liver cirrhosis is a serious symptomatic complex, caused by progressive and generally irreversible liver damage, which causes the destruction of th...